From CLARITY Wiki
(→Methodology) |
|||
| Line 1: | Line 1: | ||
| − | CLARITY is a technique developed in the Deisseroth lab at Stanford University.<ref>K Chung, J Wallace, S-Y Kim, S Kalyanasundaram, AS Andalman, TJ Davidson, JJ Mirzabekov, KA Zalocusky, J Mattis, AK Denisin, S Pak, H Bernstein, C Ramakrishnan, L Grosenick, V Gradinaru, and K Deisseroth. Structural and molecular interrogation of intact biological systems. ''Nature'' (2013) 497: 332-337.</ref><ref>K Chung and K Deisseroth. CLARITY for mapping the nervous system. ''Nature Methods'' (2013) 10(6): 508-513.</ref> The method is used to transform an intact tissue into an optically transparent and permeable hydrogel-hybridized form that can undergo immunostaining and high resolution 3-D imaging without damage to the sample. |
+ | CLARITY is a technique developed in the Deisseroth lab at Stanford University.<ref>K Chung, J Wallace, S-Y Kim, S Kalyanasundaram, AS Andalman, TJ Davidson, JJ Mirzabekov, KA Zalocusky, J Mattis, AK Denisin, S Pak, H Bernstein, C Ramakrishnan, L Grosenick, V Gradinaru, and K Deisseroth. Structural and molecular interrogation of intact biological systems. ''Nature'' (2013) 497: 332-337.</ref><ref>K Chung and K Deisseroth. CLARITY for mapping the nervous system. ''Nature Methods'' (2013) 10(6): 508-513.</ref> The method is used to transform an intact tissue into an optically transparent and permeable hydrogel-hybridized form that can undergo immunostaining and high resolution 3-D imaging without damage to the sample. By |
[[File: CLARITY_processing.png|frame|CLARITY processing on a whole mouse brain]] |
[[File: CLARITY_processing.png|frame|CLARITY processing on a whole mouse brain]] |
||
Revision as of 00:26, 20 April 2014
CLARITY is a technique developed in the Deisseroth lab at Stanford University.[1][2] The method is used to transform an intact tissue into an optically transparent and permeable hydrogel-hybridized form that can undergo immunostaining and high resolution 3-D imaging without damage to the sample. By
Contents |
Methodology
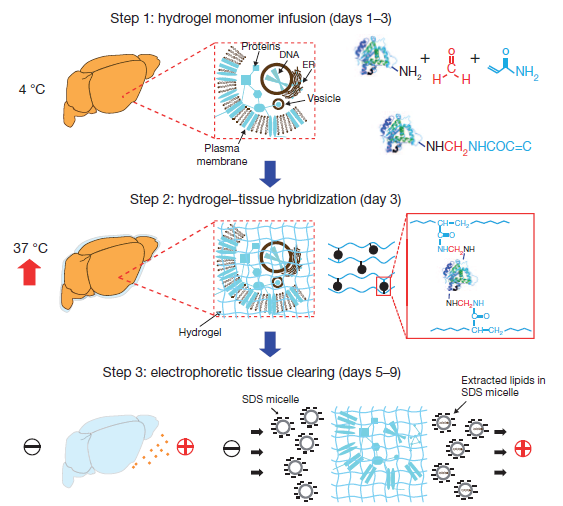
Biomolecules within the tissue are covalently linked to acrylamide monomers via reaction with formaldehyde. The acrylamide is thermally polymerized to form a hydrogel network crosslinked by the attached biomolecules and a small chemical crosslinker. An electric field is applied to actively transport SDS micelles through the tissue where they collect and remove the unattached lipids, leaving only the crosslinked biomolecules inside a swollen hydrogel matrix.
Advantages and Disadvantages
Advantages
- No damage or thin sectioning required to visualize whole intact tissue samples
- Allows marking and visualization of long-range projections and subcellular structures
- Allows multiple rounds of molecular phenotyping
Disadvantages
- Multiple-step process that takes place over several days/weeks
- Immunostaining is time-consuming for thicker tissue samples
- High start-up and consumable material costs
Tissue Samples
CLARITY can be performed on a variety of tissue types. Challenges may arise from tissues that are more fibrous (limited porosity). Tissues that contain light-scattering elements other than lipids, such as pigments, may require additional clearing mechanisms to remove those elements.
Tissues that have been successfully cleared using CLARITY include:
- Mouse brain
- Human brain slices
- Mouse spinal cord[3]
Comparison to Other Clearing Techniques
References
- ↑ K Chung, J Wallace, S-Y Kim, S Kalyanasundaram, AS Andalman, TJ Davidson, JJ Mirzabekov, KA Zalocusky, J Mattis, AK Denisin, S Pak, H Bernstein, C Ramakrishnan, L Grosenick, V Gradinaru, and K Deisseroth. Structural and molecular interrogation of intact biological systems. Nature (2013) 497: 332-337.
- ↑ K Chung and K Deisseroth. CLARITY for mapping the nervous system. Nature Methods (2013) 10(6): 508-513.
- ↑ M-D Zhang, G Tortoriello, B Hsueh, R Tomer, L Ye, N Mitsios, L Borgius, G Grant, O Kiehn, M Watanabe, M Uhlen, J Mulder, K Deisseroth, T Harkany, and TGM Hokfelt. Neuronal calcium-binding proteins 1/2 localize to dorsal root ganglia and excitatory spinal neurons and are regulated by nerve injury. PNAS (2014) 111(12): 1149-1158.